In this article we discuss how to evaluate classification accuracy, with a focus on binary classification and using software from R.
Motivation: Medical Diagnosis
Let’s say our classification is medical diagnosis, whether by a doctor of a computer algorithm. How can we decide whether the overall diagnosis ability of a doctor or computer algorithm is good? One way is to look at the accuracy, which is the (total classified correctly)/(total number of samples). However, between a false positive and a false negative for diagnosing a condition, one (often false negative) can be much worse.
For example. Consider diabetes. With
- false positive diagnosis: one may get more screening or drugs. It may also cause one to exercise more and/or eat better,.
- false negative diagnosis: one may not get treatment that is needed. This can cause the disease to worsen, leading to severe cases of diabetes.
Clearly false negatives are worse than false positives in this example, and thus it’s important to differentiate between them rather than looking solely at accuracy.
Classification Categories and the Confusion Matrix
There are four major categories in binary classification.
- True positive
- True negative
- False positive
- False negative
One can make a confusion matrix to summarize them
| Positive True Value | Negative True Value | |
| Positive Prediction | True positive | False positive |
| Negative Prediction | False negative | True negative |
Let’s load a dataset (Pima Indians Diabetes Dataset) [1], fit a naive logistic regression model, and create a confusion matrix. First we load the data and fit the model on a 75% training split. One can likely do better by feature selection and transformations but we won’t worry about that for now.
library(caret)
library(e1071)
library(pROC)
diabetes<-read.csv(url('https://raw.githubusercontent.com/jbrownlee/Datasets/master/pima-indians-diabetes.data.csv'))
N_all = nrow(diabetes)
N_train = round(0.75*(N_all))
N_test = N_all-N_train
diabetes_train <- diabetes[1:N_train,]
diabetes_test <- diabetes[N_train+1:N_all,]
classifier <- glm(X1~X6+X148+X72+X35+X0+X33.6+X0.627+X50,family='binomial',data=diabetes_train)
Then we make predictions and display the confusion matrix.
predictions <- predict(classifier,newdata=diabetes_test,type='response')[1:N_test]
confusionMatrix(factor(round(predictions)),factor(diabetes_test['X1'][1:N_test,]))
Confusion Matrix and Statistics
Reference
Prediction 0 1
0 113 31
1 9 39
Accuracy : 0.7917
95% CI : (0.7273, 0.8468)
No Information Rate : 0.6354
P-Value [Acc > NIR] : 2.057e-06
Kappa : 0.5181
Mcnemar's Test P-Value : 0.0008989
Sensitivity : 0.9262
Specificity : 0.5571
Pos Pred Value : 0.7847
Neg Pred Value : 0.8125
Prevalence : 0.6354
Detection Rate : 0.5885
Detection Prevalence : 0.7500
Balanced Accuracy : 0.7417
'Positive' Class : 0
Here we see that the accuracy is 79%, but the confusion matrix also gives interesting information. The true positive is high relative to both the false positive and false negative, while the true negative is not high relative to the false positive. We will go into more detail on some of the summaries given in the printout above in the next sections. The eight ratios right above ‘Positive’ Class at the bottom are ratios taken by dividing a table entry by a sum over a row or column.
Sensitivity and Specificity
In medical settings, sensitivity and specificity are the two most reported ratios from the confusion matrix. They are
- sensitivity: true positive rate (true positive)/(true positive+false negative). This describes what proportion of patients with diabetes are correctly identified as having diabetes. If high, we aren’t missing many people with diabetes. If low, we are, and they won’t receive the treatment they should.
- specificity: true negative rate (true negative)/(true negative+false positive). What proportion of healthy patients are correctly identified as healthy? If high, we are marking healthy as healthy. If low, we have false positives and people will either incorrectly receive treatment or in some other way incorrectly respond to a false positive.
In our diabetes example, we had a sensitivity of 0.9262. Thus if this classifier predicts that one doesn’t have diabetes, one probably doesn’t. On the other hand specificity is 0.5571429. Thus if the classifiers says that a patient has diabetes, there is a good chance that they are actually healthy.
The Receiver Operating Characteristic Curve
An important way to visualize sensitivity and specificity is via the receiving operator characteristic curve. Let’s see how we can generate this curve in R. The pROC package’s roc function is nice in that it lets one plot confidence intervals for the curve.
diabetes_sensitivity <- sensitivity(factor(round(predictions)),factor(diabetes_test['X1'][1:N_test,]))
diabetes_specificity <- specificity(factor(round(predictions)),factor(diabetes_test['X1'][1:N_test,]))
diabetes_roc <- roc(round(predictions),diabetes_test['X1'][1:N_test,],ci=TRUE,plot=TRUE, auc.polygon=TRUE, max.auc.polygon=TRUE, grid=TRUE, print.auc=TRUE, show.thres=TRUE)
diabetes.ci <- ci.se(diabetes_roc)
plot(diabetes.ci,type='shape')
plot(diabetes.ci,type='bars')
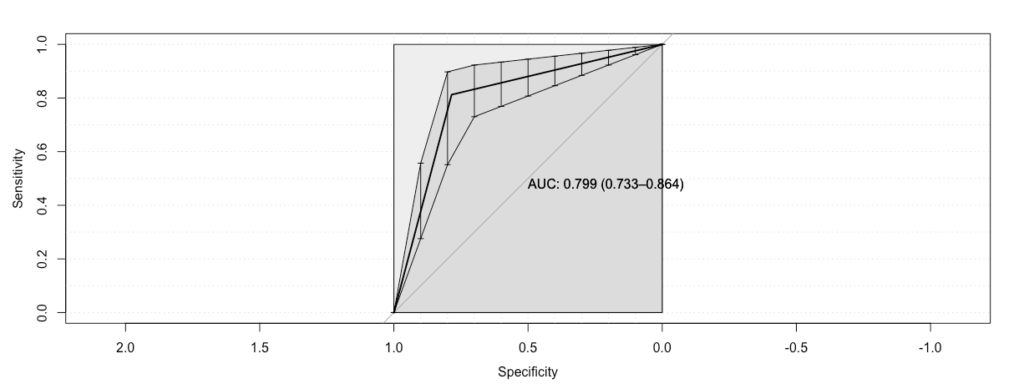
On the x-axis is specificity decreasing, and we note that specificity decreases as the false positives increase. On the y-axis is sensitivity, which increases with false positives. One interpretation of this is in terms of how much you ‘pay’ in terms of false positives to obtain true positives. The area under the curve summarizes this: if it is high you pay very little, while if it is low you pay a lot. The ‘ideal’ curve achieves sensitivity 1 for specificity 1, and has AUC 1. This implies you pay nothing in false positives for true positives. Our observed curve is pretty good though, as it has a large slope early on, and a high AUC of 0.8.
Precision, Recall, and the F1 Score
Another way to evaluate classifier accuracy, which is less common in medical diagnosis, is precision and recall. Recall is the same as sensitivity. Precision or positive predicted value is (true positives)/(true positives+false positives). In the diagnosis example, it is what proportion of diagnosed patients actually have the disease. If it is high then most of them do, while if it low then we have many false positives. For the above we had precision of 0.79, so if the prediction is positive, then there is an empirically 0.79 chance that the person actually has diabetes.
We can also combine precision and recall into an F1 score. This is the harmonic mean of precision and recall.
(1) ![]()
Conclusion
Evaluating the details of classification accuracy is important, as often the types of mistakes made by a classifier are not equally good or bad. One can do this by looking at the confusion matrix and its summaries, including precision and recall, and looking at the ROC curve and the area under the curve.
[1] Dua, D. and Graff, C. (2019). UCI Machine Learning Repository [http://archive.ics.uci.edu/ml]. Irvine, CA: University of California, School of Information and Computer Science.
But in the help, R Documentation, confusionMatrix {caret}:
Sensitivity = A / (A + C) = TP / (TP + FP)
Specificity = D / (B + D) = TN / (TN + FN)
Precision = A / (A + B) = TP / (TP + FN)
Recall = A / (A + C) = TP / (TP + FP)
I wonder why?